RNA-seqリードのシミュレーションは、バイオインフォマティクスツールの評価、比較、ベンチマーク、開発において極めて重要である。しかし、RNA-seqシミュレータの分野は過去10年間ほとんど進歩していない。このニーズに応えるため、本著者らは柔軟で高度に設定可能な設計と、ライブラリー調製とシーケンスパイプライン全体の詳細なシミュレーションを組み合わせたBEERS2を開発した。BEERS2は、カスタマイズ可能な入力またはCAMPAREEでシミュレーションされたRNAサンプルから、入力転写産物(通常、polyAテールを持つ全長のメッセンジャーRNA転写産物)を取り込む。BEERS2は、これらの転写産物のリアルなリードをFASTQ、SAMまたはBAMフォーマットで生成する。また、真の転写産物レベルの定量値も生成する。BEERS2は、柔軟で高度なコンフィギュレーションが可能な設計と、ライブラリー調製とシーケンスパイプライン全体の詳細なシミュレーションを兼ね備えており、ribosomal depletionのためのpolyA選択とRiboZeroの効果、ヘキサマープライミング配列のバイアス、ポリメラーゼ連鎖反応(PCR)増幅におけるGC-contentのバイアス、バーコードの読み取りエラー、PCR増幅中のエラーを含むように設計されている。これらの特徴を併せ持つBEERS2は、これまでで最も完全なRNA-seqのシミュレーションである。最後に、一般的なSalmon擬似アライメントアルゴリズムに対するいくつかの設定の効果を測定することで、BEERS2の利用を実証する。
インストール
https://github.com/itmat/BEERS2
mamba create -n beers2 python=3.11 -y
conda activate beers2
pip install git+https://github.com/itmat/BEERS2
> beers -h
$ beers -h
usage: beers --configfile CONFIGFILE
beers: error: the following arguments are required: --configfile
テストラン
curl https://s3.amazonaws.com/itmat.data/BEERS2/examples/baby_mouse_example.tar.gz | tar -xz
=> baby_mouse_example/ができる
解凍して出来たbaby_mouse_example/baby.config.yamlを使う。実際の使用時にもこのyamlファイルをテンプレートとして使うように書かれている。
> cat baby.config.yaml
###### BEERS CONFIGURATION ######
# The following file configures BEERS 2.
# This file is setup to run the example 'baby' data, which is
# meant as an example and to verify that BEERS2 work properly.
# The data comes from mouse, with data stripped to several small
# genomic regions.
# For the data files necesary to use this file, please run:
# wget -c https://s3.amazonaws.com/itmat.data/BEERS2/examples/baby_mouse_example.tar.gz -O - | tar -xz
# cd baby_mouse_example/
# The seed value initializes the random number generator
# so that runs are reproducible. Any integer works.
seed: 100
output:
# Output types, set to True to generate these data.
# The results appear in the 'results' directory
# FastQ Files are named as S1_L8_R2.fastq for example
# for sample 1, lane 8, read 2.
# Moreover, an 'unidentified' file is produced for each sample
# that contains reads that originated from that sample but whose
# demultiplexing step failed due to errors in the barcode reads.
output_fastq: true
# Set to true for sam/bam files containing the true alignments to
# the reference genome. Note that such alignments may no longer
# be replicable even in theory from just the reads, for exampe if
# most or all of the read is a PolyA tail and so cannot be aligned.
# This file still indicates where it came from
# Output is in the 'results' directory and named like S1_L8.sam
# for sample 1, lane 8. Like fastq, also has the potential for
# unidentified reads, stored in separate files.
output_sam: true
output_bam: false
# Set to true to receive full logs of intermediate steps
# These can be very large for a full-sized BEERS run (100s of GB)
# since they document every intermediate molecule throughout library prep
full_logs: false
global_config:
# global configuration contains configuration values that are available to all
# steps of the Library Prep and Sequencing Pipeline.
samples:
# Specify sample-specific information
# Samples are given identfiers, which are strings like '1' or '2'
'1':
# Sample one configuration
barcodes:
# Barcodes are used by AdapterLigationStep
# Each sample should have unique (i5, i7) pair of barcodes
# These get sequenced to determine which sample the read came from
# Typical examples of these can be founda at:
# https://support-docs.illumina.com/SHARE/AdapterSeq/Content/SHARE/AdapterSeq/TruSeq/UDIndexes.htm
i5: AGCGCTAG
i7: AACCGCGG
'2':
# Sample 2 configuration, same as above
barcodes:
i5: GATATCGA
i7: TTATAACC
molecule_maker_parameters:
# BEERS has two different methods for generating molecules
# Either they can be provided molecule-by-molecule in the molecule file
# Or they can be generated on demand using CAMPAREE output sample directories
# that contain all the information to synthesize random molecules
# These options affect this second on-the-fly molecules.
# The range of polyA tails to generate. Selected uniformly within this range.
min_polyA_tail_length: 50
max_polyA_tail_length: 250
resources:
# Resoruces contain general-use information
# Adapters are added to the molecules in the AdapterLigationStep
# They are also read in the sequencing by synthesis step where they
# are used to initiate transcription.
# Each adapter flanks the corresponding sample barcode, which all flanks the original molecule
# So the molecule ends up looking like (5' to 3'):
# (pre_i5_adapter) (i5) (post_i5_adapter) (molecule sequence) (pre_i7_adapter) (i7) (post_i7_adapter)
# These ones have been obtained from:
# https://support-docs.illumina.com/SHARE/AdapterSeq/Content/SHARE/AdapterSeq/TruSeq/UDIndexes.htm
pre_i5_adapter: AATGATACGGCGACCACCGAGATCTACAC
post_i5_adapter: ACACTCTTTCCCTACACGACGCTCTTCCGATCT
pre_i7_adapter: GATCGGAAGAGCACACGTCTGAACTCCAGTCAC
post_i7_adapter: ATCTCGTATGCCGTCTTCTGCTTG
# Reference genome FASTA is used to generate output BAM files, which require chromosome lengths.
# Use the same FASTA as the BEERS input (such as from CAMPAREE) was
# generated from.
reference_genome_fasta: input_data/baby_genome.mm10/baby_genome.mm10.oneline_seqs.fa
library_prep_pipeline:
# The library prep pipeline takes input molecules and walks them through a library prep
# to get them ready for sequencing.
input:
# Input the Library Prep step, which is also the input to BEERS in general
# The results of this will be fed to the sequencing pipeline to generate the output
# There are TWO ways to provide input to BEERS.
# First, we can provide molecule files, such as output by CAMPAREE
# These specify exactly the molecules to prepare and sequence
# Specify these by a path to a directory containing the molecule packets
# as if generated by CAMPAREE:
#
# /my/directory/path:
# - sample1:
# molecule_file1.txt
# molecule_file2.txt
# - sample2:
# molecule_file1.txt
# molecule_file2.txt
directory_path: input_data/
# Second, we can provide CAMPAREE sample directories
# Which instead contain the information to generate new molecules
# based off a sample's variants, gene, isoform, and allele distributions
from_distribution_data:
# Specify, for each sample, what molecule data to generate (if any)
# Samples here should correspond to the 'samples' entry above
'1':
# Number of packets to generate.
# If 0, don't generate any and just use the provided molecule files
num_packets: 1
# Number of molecules to generate per packet.
# Increasing this number increases the amount of work done per job.
# Generally want to keep this under 200000 to not run out of memory.
# If memory usage errors occur, reducing this can help.
num_molecules_per_packet: 200
# Directory containing CAMPAREE output for the sample
# This data is used to generate the resulting molecules
sample_data_directory: input_data/sample1/
'2':
num_molecules_per_packet: 200
num_packets: 1
sample_data_directory: input_data/sample2/
steps:
# The 'steps' list specifies all the steps that the Library Prep Pipeline
# will run. To specify a custom step, include it here.
# Python files with the corresponding code are looked up according to
# the pattern {module_name}.{class_name} pattern, so that the
# {module_name}.py file will be checked for {class_name} class
- step_name: polya_step.PolyAStep
# PolyA selection step removes RNA that does not have an poly A tail
# of sufficient length. Enriches for mRNA.
parameters:
# Chance per base of fragmentation prior to selection
# Increasing this above 0 induces a 3' bias.
# A value of 0.001 induces a reasonably high 3' bias
breakpoint_prob_per_base: 0.0
# Probability of retention is computed as a value between
# min_retention_prob and max_retention_prob
# For every base of the polyA tail beyond min_polya_tail_length
# the probability increases linearly from min_retention_prob
# by length_retention_prob, up to a max of max_polya_tail_length.
max_retention_prob: 1.0
min_retention_prob: 0.0
min_polya_tail_length: 40
length_retention_prob: 0.05
- step_name: fragment_step.FragmentStep
# Fragmentation step breaks each molecule into pieces
parameters:
# Fragmentation methods are available.
# 'uniform' fragments at each base equally and is the default
method: uniform
# This 'rate' is the rate parameter for an exponential distribution
# which determines the time it takes until a molecule to fragments.
# Molecules which would take longer then 'runtime' to fragment, do not fragment.
# Molecules may also fragment multiple times if rate is high enough.
rate: 0.005
runtime: 1
# Since fragmentation generates many very small fragments that will
# be quickly lost in the following steps, we have the option to
# drop those fragments immediately. Setting this value can significantly
# decrease runtime and memory requirements.
min_frag_size: 20
# If method == 'beta', then the fragmentation sites can be biased within
# the molecule, according to a beta distribution.
# NOTE: using 'beta' can generate unusual coverage plots since there are
# significant edge effects around the transcript. However, it can be used
# to give a more realistic fragment distribution.
#
# If using 'beta', you must also specify the following:
#
# The parameters for the beta distribution. Set these so that
# A = B > 0 to bias towards fragmentation in the middle of the fragment,
# with larger values biasing further towards the middle.
# If A > B, then would bias towards the 5' end instead.
# beta_A: 3.0
# beta_B: 3.0
#
# And the N factor allows a non-linear fragmentation rate depending
# upon the length of the molecule. Values >1 indicate that longer molecules
# are more likely to fragment than smaller ones, biasing towards larger
# fragment sizes.
# beta_N: 2.0
- step_name: first_strand_synthesis_step.FirstStrandSynthesisStep
# First Strand Synthesis performs (hexamer) priming on the RNA
# and then generates the cDNA from this. Priming sites can be
# biased according the position probability matrix (PPM) that
# specifies the weighting of each base according to the position
# relative to the first base of the primer.
# The 5' most primer is then used to extend the DNA and create
# the cDNA, thereby potentially losing some of the 5' end of
# the molecule.
parameters:
# If 'perfect_priming' is true, then we always prime
# exactly on the 5' most end of the molecule and the
# cDNA matches perfectly the original molecule
# If true, all other parameters are ignored.
perfect_priming: false
# PPM matrix describing biases of the priming sites
# Values must sum to 1 in each base. Length of the PPM
# determines the primer lengths.
# See Kasper et all, 2010 PMID: 20395217 for example
# observation of these biases in fragment start positions.
# If no bias is desired, set all values to 0.25.
position_probability_matrix:
A: [0.50, 0.1, 0.40, 0.30, 0.25, 0.15]
C: [0.20, 0.5, 0.3 , 0.25, 0.25, 0.15]
G: [0.15, 0.1, 0.15, 0.25, 0.25, 0.20]
T: [0.15, 0.3, 0.15, 0.20, 0.25, 0.50]
# Rate of priming: approximate number of primers that
# will bind (if bias-free) per kilobase of the fragment
# Higher numbers will loose less of the 5' side.
primes_per_kb: 50
- step_name: second_strand_synthesis_step.SecondStrandSynthesisStep
# Second Strand synthesis copies the single-stranded cDNA from
# the First Strand Synthesis step into double-stranded cDNA.
# All parameters are same as the First Strand Synthesis step.
# If not perfect_priming, will loose some of the 5' end of the
# first strand of cDNA, which is the 3' end of the original fragment
parameters:
perfect_priming: false
position_probability_matrix:
A: [0.50, 0.1, 0.40, 0.30, 0.25, 0.15]
C: [0.20, 0.5, 0.3 , 0.25, 0.25, 0.15]
G: [0.15, 0.1, 0.15, 0.25, 0.25, 0.20]
T: [0.15, 0.3, 0.15, 0.20, 0.25, 0.50]
primes_per_kb: 50
- step_name: sizing_step.SizingStep
# Sizing step throws out molecules that do not fit the
# desired size distribution
parameters:
# Probability of retention by length
# __________________
# 1| ___/ \___
# p | ____/ \____
# 0|___/ \____
# -----------------------------------------------
# | | | |
# min_length | | max_length
# | select_all |
# start end
#
# All molecules with lengths outside of this range
# will be discarded:
min_length: 100
max_length: 400
# Molecules whose length is inside this range will
# always be retained. Retention probability raises
# linearly from 0 to 1 between the min/max values
# above and this range here.
select_all_start_length: 200
select_all_end_length: 300
- step_name: adapter_ligation_step.AdapterLigationStep
# Adpater Ligation attaches adapters to each end of the molecules
# These are used for the PCR step to initiate PCR and include the
# sample identifying barcodes.
#
# This uses the adapters specified in the resources section
# as well as the sample i5/i7 barcodes in the samples section
parameters: {}
- step_name: pcr_amplification_step.PCRAmplificationStep
# PCR amplification step creates many copies of each molecule
# Since the number of molecules generated is quite large and
# since in real sequencing, only a small fraction of the molecules
# prepped end up forming clusters on the flowcell and being
# sequenced, we give the option to remove most of the PCR
# generated molecules prior to the end of the step.
# This provides a very large increase in speed and decrease
# in memory usage.
parameters:
# Number of cycles to use
# Generates 2^n fragments for each input fragment
number_cycles: 10
# Retain only this percent of the data
# NOTE: scale is 0-100, not 0-1, so 0.08 means 0.0008 of the generated molecules are kep
# This value can be approximated from real data with UMI tags by the following:
# Compute the PCR duplication rate using the UMI tags.
# Let N be the number of PCR steps. Assuming that each molecule generates 2^N
# PCR descendants, if all were sequenced, we would expect all molecules to be duped
# 2^N times. Instead, choose retention_percentage with the following:
#
# '''
# import scipy.stats
# pcr_rate = scipy.stats.binom(p=retention_percentage / 100, n=2^N).sf(1)
# # Choose retention_percentage so that pcr_rate is approximately the observed PCR rate
# '''
#
# For example, retention_percentage = 0.08 and number_cycles=10 gives dupe rate
# of about 20%, which is pretty typical. If number_cycles changes, this should
# be modified too to maintain dupe rate.
retention_percentage: 0.08
# Induce a GC bias by discarding some PCR duplicates
# according to their GC bias. All fragments overall
# GC content is computed and then the following three
# parameters are used to compute a probability of retention
# gc_bias_constant + gc_bias_linear*(gc - 0.5) + gc_bias_quadratic*(gc - 0.5)^2
# clipped to always be within 0 and 1
# For no bias, set to 1, 0, and 0 for const, linear, and quadratic, respectively.
# To bias towards GC=0.5 content, set gc_bias_quadratic to be negative,
# such as -100 for a large GC bias.
gc_bias_constant: 1.0
gc_bias_linear: 0.0
gc_bias_quadratic: -100
# During PCR, we have some chance of mis-copying the molecules
# These are specified here as probability per-base, so 0.001 is one error per 1kb
deletion_rate: 0.0001
insertion_rate: 0.0001
substitution_rate: 0.001
sequence_pipeline:
# The Sequence Pipeline takes clusters bound to the flowcell and turns them into
# final, sequenced results, including fastq files or SAM files with the 'true'
# alignments of the simulated molecules to the reference genome.
steps:
- step_name: bridge_amplification_step.BridgeAmplificationStep
# Bridge amplification takes a seed of a cluster and generates
# many copies of the molecule to form a cluster.
parameters:
# Number of cycles to perform. Generates 2^N molecules in the cluster
# via a PCR-like bridge amplification process.
cycles: 10
# Substitution rate per base in the cluster formation
# No indels are generated here, for computation simplicity.
substitution_rate: 0.01
- step_name: sequence_by_synthesis_step.SequenceBySynthesisStep
# Sequencing by Synthesis then synthesizes complementary strands to each
# molecule in the clusters with flourescence readings at each base.
# If some of the molecules have errors in them (from bridge amplification),
# the flourescence will be imperfect. Moreover, errors also happen in this step
# including phasing, where some molecules get ahead or behind in the sequencing
# and so are displaying the wrong base.
# From these readings, the final sequenced values are generated.
parameters:
# Determines which read is the forward and which is the reverse read
# NOTE: currently only 'true' is implemented
forward_is_5_prime: true
# Whether to sequence both ends or just one
# NOTE: currently only 'true' is implemented
paired_ends: true
# Length of the reads to generate, in bases
read_length: 100
# The rate of phasing, either forward (skip) or backwards (drop)
# per base per molecule
skip_rate: 0.002
drop_rate: 0.002
flowcell:
# BEERS simulates one flowcell
# Layout of the flowcell is defined here
# Coordinate strategy defines how coordinates are assigned
# Currently only 'random' is supported
coordinate_strategy: random
# Flowcell geometry defines the dimensions of the flowcell
# Example is given for a typical HiSeq 2500 flowcell
flowcell_geometry:
# Range of the lane numbers
min_lane: 1
max_lane: 8
# Range of the tile values
max_tile: 2228
min_tile: 1101
# Range of the x-coordinates
min_x: 1012
max_x: 32816
# Range of the y-coordinates
min_y: 998
max_y: 49247
# List of lanes that samples will be written into
# Molecules are distributed evenly across lanes
# These must all be within [min_lane, max_lane] range
lanes_to_use: [1, 2]
パラメータが決まったら実行する。
cd baby_mouse_example/
#BEERS2のラン
beers --configfile baby.config.yaml --jobs 1
テスト時は1分以内に終了した。
出力例
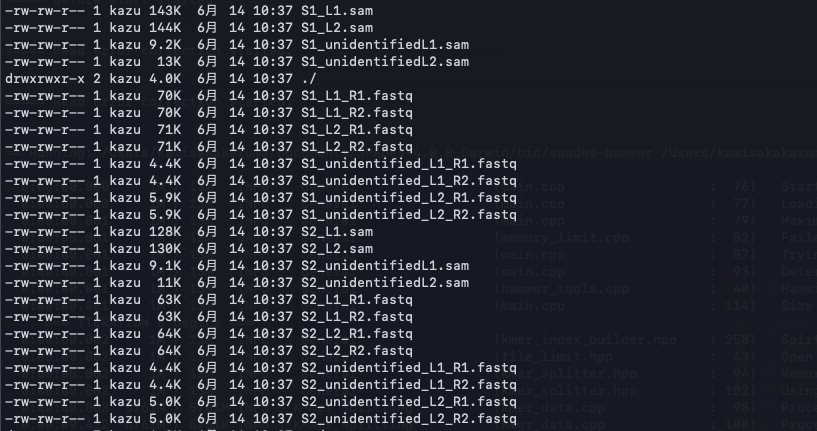
デフォルトでは、fastqの他にゴールデンスタンダードなsamも出力される("output_sam: true"の部分)。
引用
BEERS2: RNA-Seq simulation through high fidelity in silico modeling
Thomas G Brooks, Nicholas F Lahens, Antonijo Mrčela, Dimitra Sarantopoulou, Soumyashant Nayak, Amruta Naik, Shaon Sengupta, Peter S Choi, Gregory R Grant
Briefings in Bioinformatics, Volume 25, Issue 3, May 2024