2019 2/15 Biocondaインストール追加、バッチモード追加
2019 3/10 タイトル修正
2019 5/50 インストール方法追記
ARAGORNは既存のtRNAとのホモロジーや二次構造などを手掛かりにゲノム中からtRNAやtmRNAを探すツール。
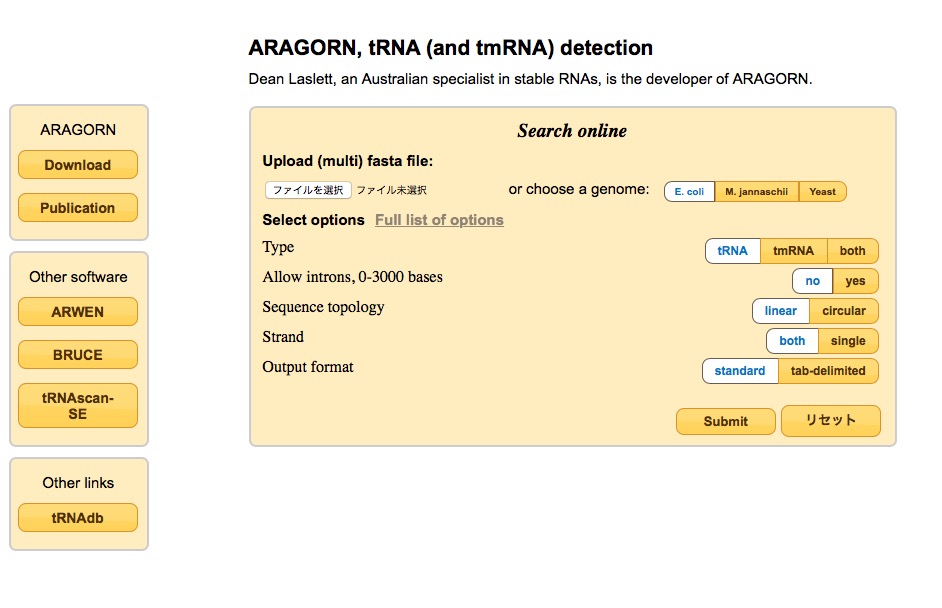
webサーバー
ARAGORN, tRNA (and tmRNA) detection
インストール
brewで導入できる。
#biocoonda
conda install -c bioconda -y ARAGORN
#homebrew
brew install ARAGORN
実行方法
aragorn genome.fa -o tRNAs_output
ポジションごとにtRNAが予測される。

最後にまとめが表示される。

例えばここで使ったゲノムではバクテリアのクロモソームから45 tRNAが見つかり、プラスミドからはtRNAは見つからなかった。tRNAのアンチコドンの頻度も表示されている。
genetic codeがスタンダードかどうかなどはフラグを立てて設定する。ARAGORN -hで確認できるフラグの一覧を貼っておく。
user$ ARAGORN -h
----------------------------
ARAGORN v1.2.36 Dean Laslett
----------------------------
Please reference the following papers if you use this
program as part of any published research.
Laslett, D. and Canback, B. (2004) ARAGORN, a
program for the detection of transfer RNA and transfer-messenger
RNA genes in nucleotide sequences
Nucleic Acids Research, 32;11-16
Laslett, D. and Canback, B. (2008) ARWEN: a
program to detect tRNA genes in metazoan mitochondrial
nucleotide sequences
Bioinformatics, 24(2); 172-175.
ARAGORN detects tRNA, mtRNA, and tmRNA genes.
Usage:
aragorn -v -s -d -c -l -a -w -j -ifro<min>,<max> -t -mt -m -tv -gc -seq -br -fasta -fo -o <outfile> <filename>
<filename> is assumed to contain one or more sequences
in FASTA format. Results of the search are printed to
STDOUT. All switches are optional and case-insensitive.
Unless -i is specified, tRNA genes containing introns
are not detected.
-m Search for tmRNA genes.
-t Search for tRNA genes.
By default, both are detected. If one of
-m or -t is specified, then the other
is not detected unless specified as well.
-mt Search for Metazoan mitochondrial tRNA
genes. -i switch ignored. Composite
Metazoan mitochondrial genetic code used.
-mtmam Search for Mammalian mitochondrial tRNA
genes. -i switch ignored. -tv switch set.
Mammalian mitochondrial genetic code used.
-mtx Same as -mt but low scoring tRNA genes are
not reported.
-gc<num> Use the GenBank transl_table = <num> genetic code.
-gcstd Use standard genetic code.
-gcmet Use composite Metazoan mitochondrial genetic code.
-gcvert Use Vertebrate mitochondrial genetic code.
-gcinvert Use Invertebrate mitochondrial genetic code.
-gcyeast Use Yeast mitochondrial genetic code.
-gcprot Use Mold/Protozoan/Coelenterate mitochondrial genetic code.
-gcciliate Use Ciliate genetic code.
-gcflatworm Use Echinoderm/Flatworm mitochondrial genetic code.
-gceuplot Use Euplotid genetic code.
-gcbact Use Bacterial/Plant Chloroplast genetic code.
-gcaltyeast Use alternative Yeast genetic code.
-gcascid Use Ascidian Mitochondrial genetic code.
-gcaltflat Use alternative Flatworm Mitochondrial genetic code.
-gcblep Use Blepharisma genetic code.
-gcchloroph Use Chlorophycean Mitochondrial genetic code.
-gctrem Use Trematode Mitochondrial genetic code.
-gcscen Use Scenedesmus obliquus Mitochondrial genetic code.
-gcthraust Use Thraustochytrium Mitochondrial genetic code.
Individual modifications can be appended using
,BBB=<aa> B = A,C,G, or T. <aa> is the three letter
code for an amino-acid. More than one modification
can be specified. eg -gcvert,aga=Trp,agg=Trp uses
the Vertebrate Mitochondrial code and the codons
AGA and AGG changed to Tryptophan.
-tv Do not search for mitochondrial TV replacement
loop tRNA genes. Only relevant if -mt used.
-i Search for tRNA genes with introns in
anticodon loop with maximum length 3000
bases. Minimum intron length is 0 bases.
Ignored if -m is specified.
-i<max> Search for tRNA genes with introns in
anticodon loop with maximum length <max>
bases. Minimum intron length is 0 bases.
Ignored if -m is specified.
-i<min>,<max> Search for tRNA genes with introns in
anticodon loop with maximum length <max>
bases, and minimum length <min> bases.
Ignored if -m is specified.
-io Same as -i, but allow tRNA genes with long
introns to overlap shorter tRNA genes.
-if Same as -i, but fix intron between positions
37 and 38 on C-loop (one base after anticodon).
-ifo Same as -if and -io combined.
-ir Same as -i, but search for tRNA genes with minimum intron
length 0 bases, and only report tRNA genes with minimum
intron length <min> bases.
-c Assume that each sequence has a circular
topology. Search wraps around each end.
Default setting.
-l Assume that each sequence has a linear
topology. Search does not wrap.
-d Double. Search both strands of each
sequence. Default setting.
-s or -s+ Single. Do not search the complementary
(antisense) strand of each sequence.
-sc or -s- Single complementary. Do not search the sense
strand of each sequence.
-ss Use the stricter canonical 1-2 bp spacer1 and
1 bp spacer2. Ignored if -mt set. Default is to
allow 3 bp spacer1 and 0-2 bp spacer2, which may
degrade selectivity.
-ps Lower scoring thresholds to 95% of default levels.
-ps<num> Change scoring thresholds to <num> percent of default levels.
-rp Flag possible pseudogenes (score < 100 or tRNA anticodon
loop <> 7 bases long). Note that genes with score < 100
will not be detected or flagged if scoring thresholds are not
also changed to below 100% (see -ps switch).
-seq Print out primary sequence.
-br Show secondary structure of tRNA gene primary
sequence with round brackets.
-fasta Print out primary sequence in fasta format.
-fo Print out primary sequence in fasta format only
(no secondary structure).
-fon Same as -fo, with sequence and gene numbering in header.
-fos Same as -fo, with no spaces in header.
-fons Same as -fo, with sequence and gene numbering, but no spaces.
-j Display 4-base sequence on 3' end of astem
regardless of predicted amino-acyl acceptor
length.
-jr Allow some divergence of 3' amino-acyl acceptor
sequence from NCCA.
-jr4 Allow some divergence of 3' amino-acyl acceptor
sequence from NCCA, and display 4 bases.
-v Verbose. Prints out search progress
to STDERR.
-a Print out tRNA domain for tmRNA genes
-o <outfile> print output into <outfile>. If <outfile>
exists, it is overwritten.
By default, output goes to STDOUT.
-w Print out genes in batch mode.
For tRNA genes, output is in the form:
Sequence name
N genes found
1 tRNA-<species> [locus 1] <Apos> (nnn)
i(<intron position>,<intron length>)
.
.
N tRNA-<species> [Locus N] <Apos> (nnn)
i(<intron position>,<intron length>)
N is the number of genes found
<species> is the tRNA iso-acceptor species
<Apos> is the tRNA anticodon relative position
(nnn) is the tRNA anticodon base triplet
i means the tRNA gene has a C-loop intron
For tmRNA genes, output is in the form:
n tmRNA(p) [Locus n] <tag offset>,<tag end offset>
<tag peptide>
p means the tmRNA gene is permuted
リストを作るならバッチモードでランする
ARAGORN genome.fa -o output -w
>NODE_1
38 genes found
1 tRNA-Phe c[27015,27088] 35 (gaa)
2 tRNA-Val c[27566,27640] 35 (tac)
3 tRNA-Arg c[41028,41100] 35 (gcg)
4 tRNA-Asn [46340,46415] 35 (gtt)
5 tRNA-Met [46592,46666] 36 (cat)
6 tRNA-Arg [145784,145858] 35 (ccg)
7 tRNA-Arg c[255331,255424] 35 (tcg)
8 tRNA-Tyr [289892,289966] 35 (gta)
9 tRNA-Gln [469893,469965] 35 (ttg)
10 tRNA-Ser [534698,534785] 28 (gct)
11 tRNA-Ser [585265,585349] 36 (cga)
12 tRNA-Gly c[699904,699975] 34 (gcc)
13 tRNA-Val c[752912,752986] 36 (gac)
14 tRNA-Val c[802618,802696] 37 (cac)
15 tRNA-Ala [841719,841790] 33 (ggc)
16 tRNA-Thr [882862,882935] 35 (cgt)
17 tRNA-Gly c[920091,920169] 38 (gcc)
18 tRNA-Arg c[921771,921847] 37 (tcg)
19 tRNA-Thr c[922371,922444] 35 (ggt)
20 tRNA-His c[922446,922519] 35 (gtg)
21 tRNA-Glu c[923590,923666] 37 (ttc)
22 tRNA-Cys c[923671,923744] 35 (gca)
23 tRNA-Asn c[923783,923858] 36 (gtt)
24 tRNA-Lys c[923936,924011] 36 (ttt)
25 tRNA-Tyr c[924144,924216] 35 (gta)
26 tRNA-Ala c[924221,924294] 34 (cgc)
27 tRNA-Met c[924317,924391] 37 (cat)
28 tRNA-Leu c[924394,924476] 34 (cag)
29 tRNA-Leu c[924509,924590] 35 (gag)
30 tRNA-Leu c[924593,924676] 36 (taa)
31 tRNA-Trp c[924678,924748] 33 (cca)
32 tRNA-Ser c[924751,924838] 38 (gga)
33 tRNA-Ser c[924839,924922] 36 (tga)
34 tRNA-Ser c[924957,925046] 36 (gct)
35 tRNA-Pro c[925049,925121] 35 (tgg)
36 tRNA-Asp c[925322,925396] 36 (gtc)
37 tRNA-Gln c[925408,925483] 37 (ttg)
38 tRNA-Arg c[927017,927091] 36 (tcg)
引用
ARAGORN, a program to detect tRNA genes and tmRNA genes in nucleotide sequences
Dean Laslett Bjorn Canback
Nucleic Acids Research, Volume 32, Issue 1, 1 January 2004, Pages 11–16,